Chapter 5: Q34P. (page 129)
You wish to sequence the light chain of a protease inhibitor from the Brassica nigra plant. Cleavage of the light chain by trypsin and chymotrypsin yields the following fragments. What is the sequence of the light chain?
Chymotrypsin
1. Leu–His–Lys–Gln–Ala–Asn–Gln–Ser–Gly–Gly–Gly–Pro–Ser
2. Gln–Gln–Ala–Gln–His–Leu–Arg–Ala–Cys–Gln–Gln–Trp
3. Arg–Ile–Pro–Lys–Cys–Arg–Lys–Phe
Trypsin
4. Arg
5. Ala–Cys–Gln–Gln–Trp–Leu–His–Lys
6. Cys–Arg
7. Gln–Ala–Asn–Gln–Ser–Gly–Gly–Gly–Pro–Ser
8. Phe–Gln–Gln–Ala–Gln–His–Leu–Arg
9. Ile–Pro–Lys
10. Lys
Short Answer
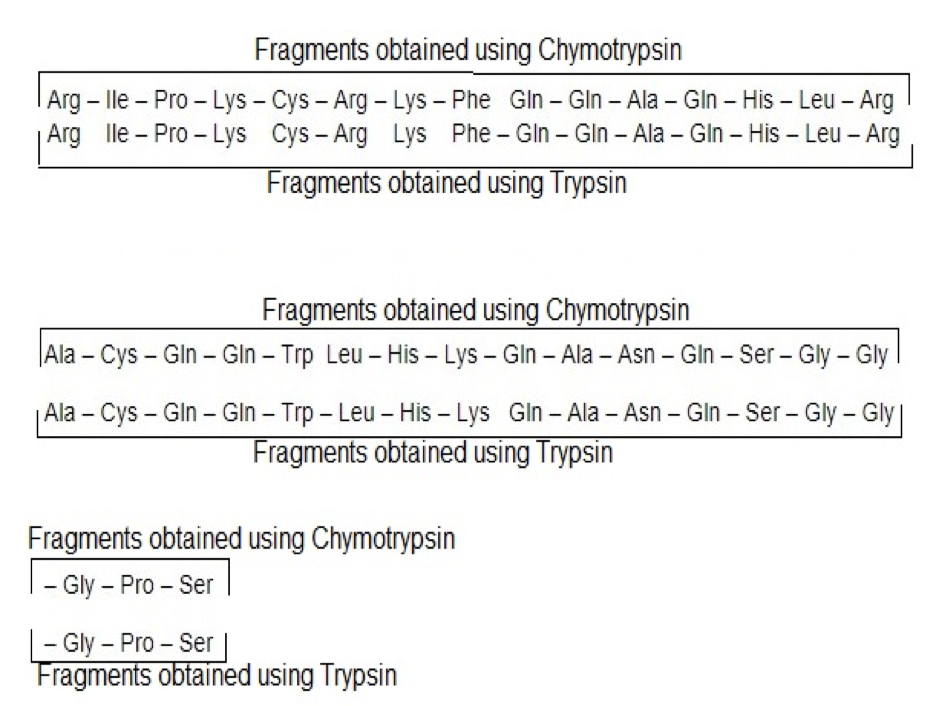
The light chain will have the sequence:
Arg – Ile – Pro – Lys – Cys – Arg – Lys – Phe – Gln – Gln – Ala – Gln – His – Leu – Arg – Ala – Cys – Gln – Gln – Trp Leu – His – Lys – Gln – Ala – Asn – Gln – Ser – Gly – Gly – Gly – Pro – Ser.