Chapter 24: Q-24-47P (page 1300)
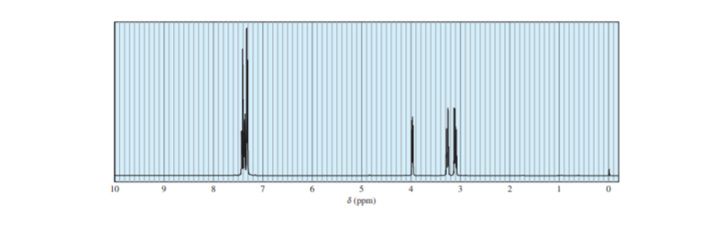
A student took the proton NMR spectrum of phenylalanine in D2Osolution, and had the instrument suppress the DOHsolvent peak. The spectrum is shown below. The integrated relative areas of the peaks are 5:1:1:1.
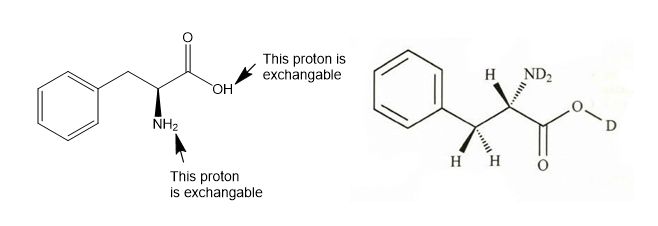
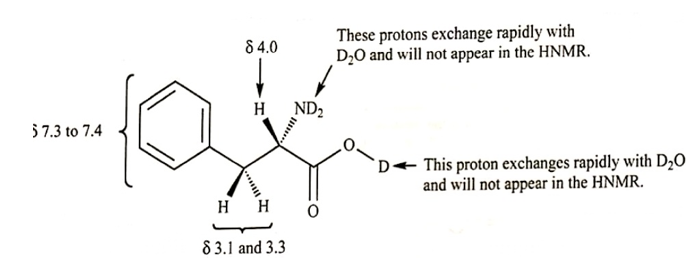
(a) Draw the structure of phenylalanine as it exists in D2Osolution. (There is a large excess of D2O, and any exchangeable protons in phenylalanine will exchange with the solvent.)
(b) Assign the peaks in the spectrum to the protons in the structure.
(c) Why don’t we see the -NH2or -COOHprotons in the spectrum?
(d) What is the relationship between the two protons that generate nearly mirror-image multiplets at 3.1 and 3.3?
Short Answer
(a)
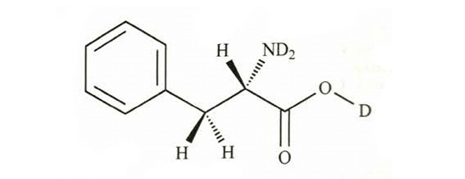
(b)
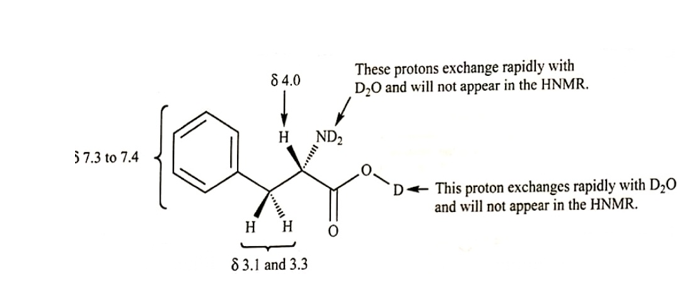
(c) In D2O, there is rapid exchange between protons on carboxylic group and amino group, thus, when H is replaced with D, no signal gets generated in NMR.
(d) The relation between two protons that generate nearly mirror-image multiplets is diastereotopic in nature.