Fluorescence quenching in micelles. Consider an aqueous solution with a high concentration of micelles and relatively low concentrations of the fluorescent molecule pyrene and a quencher (cetylpyridinium chloride, designated Q), both of which dissolve in the micelles.
Quenching occurs if pyrene and Q are in the same micelle. Let the total concentration of quencher be [Q] and the concentration of micelles be [M]. The average number of quenchers per micelle is. If Q is randomly distributed among the micelles, then the probability that a particular micelle has n molecules of Q is given by the Poisson distribution:
Probability of n molecules of Q in micelle =
whereis n factorial. The probability that there are no molecules of Q in a micelle is
Probability ofmolecules of Q in micelle =
because 0!=1
Let be the fluorescence intensity of pyrene in the absence of Q and let IQbe the intensity in the presence of Q (both measured at the same concentration of micelles). The quotient must be which is the probability that a micelle does not possess a quencher molecule. Substituting gives
Micelles are made of the surfactant molecule, sodium dodecyl sulfate. When surfactant is added to a solution, no micelles form until a minimum concentration called the critical micelle concentration (CMC) is attained. When the total concentration of surfactant, [S], exceeds the critical concentration, then the surfactant found in micelles is. The molar concentration of micelles is
where Nav is the average number of molecules of surfactant in each micelle.
Combining Equationsandgives an expression for fluorescence as a function of total quencher concentration, [Q]:
By measuring fluorescence intensity as a function of [Q] at fixed [S], we can find the average number of molecules of S per micelle if we know the critical micelle concentration (which is independently measured in solutions of S). The table gives data for
pyrene in a micellar solution with a total concentration of sodium dodecyl sulfate [S]=20.8mM
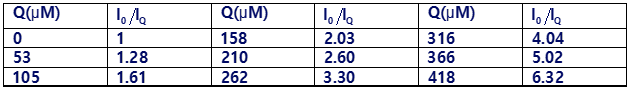
(a) If micelles were not present, quenching would be expected to follow the Stern-Volmer equation. Show that the graph of versus [Q] is not linear.
(b) The critical micelle concentration is 8.1mM.Prepare a graph ofversus [Q]. Use Equation 5 to find Nav, the average number of sodium dodecyl sulfate molecules per micelle.
(c) Find the concentration of micelles, [M], and the average number of molecules of Q per micelle,, when
(d) Compute the fractions of micelles containing,, andmolecules of Q when









