Chapter 24: Q4P (page 875)
Calculate the contour length for the gene in in Problem 3 if it assumed an A-DNA conformation.
Short Answer
The contour length for the gene if it is an A-DNA conformation is or .
Learning Materials
Features
Discover
Chapter 24: Q4P (page 875)
Calculate the contour length for the gene in in Problem 3 if it assumed an A-DNA conformation.
The contour length for the gene if it is an A-DNA conformation is or .
All the tools & learning materials you need for study success - in one app.
Get started for free
Explain why most nucleotides adopt the anti conformation.
Polyoma virus DNA can be separated by sedimentation in an ultracentrifuge (Section 5-2E) at neutral pH into three components that have sedimentation coefficients of 20, 16, and 14.5S and that are known as Types I, II, and III DNAs, respectively. These DNAs all have identical base sequences and molecular masses. In 0.15 M NaCl, both Types II and III DNA have melting curves of normal cooperativity and a Tm of 88°C. Type I DNA, however, exhibits a very broad melting curve and a Tm of 107°C. At pH 13, Types I and III DNAs have sedimentation coefficients of 53 and 16S, respectively, and Type II separates into two components with sedimentation coefficients of 16 and 18S. How do Types I, II, and III DNAs differ from one another? Explain their different physical properties.
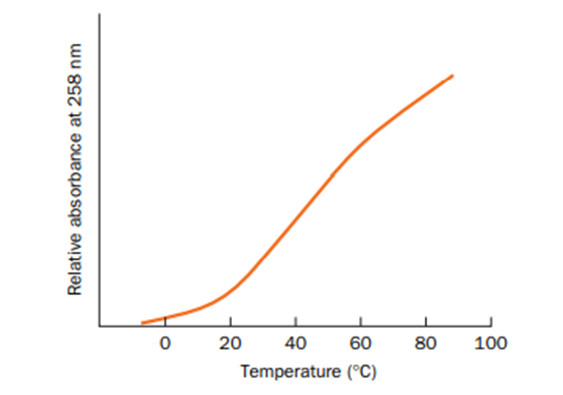
The melting curve for the polyribonucleotide poly(A) is shown below.
(a) Explain why absorbance increases with increasing temperature.
(b) Why does the shape of the curve differ from the one shown in Fig. 24-22?
Most protein-coding genes are present in just one copy per genome. Explain why eukaryotic genomes typically contain dozens of copies of histone genes.
How does the DNA binding of a zinc finger–containing transcription factor differ from that of a prokaryotic repressor?
What do you think about this solution?
We value your feedback to improve our textbook solutions.